Parts 1 and 2 made the case that equal-area cells matter for biodiversity counts. That argument is already won in practice: global biodiversity maps are routinely projected through equal-area cylindrical projections such as Behrmann or Mollweide before binning, precisely so that cell counts reflect density rather than projection geometry.
So why DGGS instead of an equal-area projection?
Because for AI-ready, multi-resolution, cloud-native analysis, equal-area is not enough. An equal-area projection fixes the area bias but introduces three new problems:
Cell shape distortion (anisotropy). A Behrmann cell at the equator is square-ish. The same cell at 70°N is the same area but several times wider than tall. A CNN’s 3×3 kernel covers the same number of cells everywhere, but a wildly different physical shape of geography. Models cannot learn consistent spatial operators on inconsistent neighborhoods.
No native multi-resolution hierarchy. Refining a 100 km Behrmann grid to 50 km requires reprojection and rebinning, with interpolation artefacts. HEALPix
nsidedoubling produces exactly 4 children per parent, deterministically, with no resampling.Pole degeneracies (Mollweide). Mollweide compresses both poles to single points; polar cells degenerate. Behrmann avoids this but pays in extreme cell elongation. HEALPix uses no projection — cells live directly on the sphere.
This notebook illustrates (1) — the most visually direct of the three arguments — and quantifies it across the latitudes that actually matter for biodiversity (0° to 70°N).
import os
import shutil
import cartopy.crs as ccrs
import geopandas as gpd
import h3
import healpix_geo.nested as hgn
import healpy as hp
import matplotlib.pyplot as plt
import numpy as np
from dggrid4py import DGGRIDv7
from matplotlib.patches import Polygon
from rhealpixdggs.rhp_wrappers import geo_to_rhp, rhp_to_geo_boundary
from shapely.geometry import Point
R = 6371.0 # Earth radius, km
DEG = np.pi / 180.0
# DGGRID handle, used by ISEA3H cell builder. Found via DGGRID_PATH env
# var (set in Docker image) or PATH (conda-forge dggrid).
_dggrid_bin = os.environ.get("DGGRID_PATH") or shutil.which("dggrid")
_dggrid = DGGRIDv7(executable=_dggrid_bin, working_dir="/tmp",
capture_logs=True, silent=True) if _dggrid_bin else NoneCell builders¶
For each grid system we build the geographic footprint (vertex lat/lon array) of a representative cell whose centroid is at a given latitude. Cells are densified along their edges so that curvature is preserved when reprojected.
def latlon_cell_vertices(lat_c, lon_c, dlat=5.0, dlon=5.0, n_edge=40):
"""5° lat-lon cell centred at (lat_c, lon_c)."""
lat_s, lat_n = lat_c - dlat / 2, lat_c + dlat / 2
lon_w, lon_e = lon_c - dlon / 2, lon_c + dlon / 2
bottom = np.linspace(lon_w, lon_e, n_edge)
right = np.linspace(lat_s, lat_n, n_edge)
top = np.linspace(lon_e, lon_w, n_edge)
left = np.linspace(lat_n, lat_s, n_edge)
lats = np.r_[np.full(n_edge, lat_s), right, np.full(n_edge, lat_n), left]
lons = np.r_[bottom, np.full(n_edge, lon_e), top, np.full(n_edge, lon_w)]
return lats, lons
def behrmann_cell_vertices(lat_c, lon_c, equator_dlat=5.0, equator_dlon=5.0, n_edge=40):
"""Behrmann (cylindrical equal-area, std parallels ±30°) cell.
Cell is constructed so that all cells have the SAME AREA as a
`equator_dlat` × `equator_dlon` cell at the equator. Holding
longitude width constant at `equator_dlon`, the latitude span at
`lat_c` solves sin(φ_n) − sin(φ_s) = sin(equator_dlat/2)·2.
"""
sin_half_target = np.sin(equator_dlat / 2 * DEG)
# sin(φ_n) − sin(φ_s) = 2·cos(φ_c)·sin(δ/2)
# → δ = 2·arcsin(sin_half_target / cos(φ_c))
cos_c = np.cos(lat_c * DEG)
arg = sin_half_target / cos_c
if abs(arg) >= 1.0:
# Cell would extend beyond the pole; clamp.
arg = np.sign(arg) * 0.9999
dlat = 2 * np.degrees(np.arcsin(arg))
return latlon_cell_vertices(
lat_c, lon_c, dlat=dlat, dlon=equator_dlon, n_edge=n_edge
)
def healpix_cell_vertices(lat_c, lon_c, nside=16, n_edge=20):
"""HEALPix cell whose centre is closest to (lat_c, lon_c)."""
theta = (90.0 - lat_c) * DEG
phi = (lon_c % 360) * DEG
pix = hp.ang2pix(nside, theta, phi, nest=True)
# boundaries() returns 3xN cartesian; convert to lat/lon
xyz = hp.boundaries(nside, pix, step=n_edge, nest=True) # (3, 4*n_edge)
x, y, z = xyz
cell_lat = 90.0 - np.degrees(np.arccos(z))
cell_lon = np.degrees(np.arctan2(y, x))
return cell_lat, cell_lon
def healpix_geo_cell_vertices(lat_c, lon_c, depth=4):
"""HEALPix cell on the WGS84 ellipsoid (via healpix-geo / cdshealpix).
`healpix-geo` implements the GRID4EARTH "Ellipsoidal HEALPix" path:
HEALPix indexing whose cell vertices are computed on the WGS84
ellipsoid via authalic-sphere mapping. depth=4 (nside=16) matches
the HEALPix-on-sphere resolution used elsewhere in this notebook.
"""
centre = hgn.lonlat_to_healpix(
np.array([float(lon_c)]), np.array([float(lat_c)]),
depth=depth, ellipsoid="WGS84",
)
lons_v, lats_v = hgn.vertices(
centre.astype(np.uint64), depth=depth, ellipsoid="WGS84",
)
return np.asarray(lats_v[0]), np.asarray(lons_v[0])
def h3_cell_vertices(lat_c, lon_c, resolution=3):
"""H3 cell containing (lat_c, lon_c). Returns (lats, lons)."""
cell = h3.latlng_to_cell(float(lat_c), float(lon_c), resolution)
boundary = h3.cell_to_boundary(cell) # tuple of (lat, lng)
lats = np.array([b[0] for b in boundary])
lons = np.array([b[1] for b in boundary])
return lats, lons
def rhealpix_cell_vertices(lat_c, lon_c, resolution=4):
"""rHEALPix cell containing (lat_c, lon_c). Returns (lats, lons).
NOTE: rhp_to_geo_boundary returns (lon, lat) — opposite order to H3.
"""
cell = geo_to_rhp(float(lat_c), float(lon_c), resolution, plane=False)
boundary = rhp_to_geo_boundary(cell, plane=False)
lats = np.array([b[1] for b in boundary])
lons = np.array([b[0] for b in boundary])
return lats, lons
def isea3h_cell_vertices(lat_c, lon_c, resolution=8):
"""ISEA3H cell containing (lat_c, lon_c) (via dggrid4py)."""
if _dggrid is None:
raise RuntimeError(
"DGGRID binary not found. Install conda-forge dggrid=8.41 "
"or run inside the Docker container."
)
pt_gdf = gpd.GeoDataFrame(
{"id": [0]},
geometry=[Point(float(lon_c), float(lat_c))],
crs="EPSG:4326",
)
cells_df = _dggrid.cells_for_geo_points(
geodf_points_wgs84=pt_gdf, cell_ids_only=True,
dggs_type="ISEA3H", resolution=resolution,
)
cell_id = int(cells_df["name"].iloc[0])
poly_gdf = _dggrid.grid_cell_polygons_from_cellids(
cell_id_list=[cell_id], dggs_type="ISEA3H", resolution=resolution,
clip_subset_type="WHOLE_EARTH",
)
geom = poly_gdf.geometry.iloc[0]
coords = list(geom.exterior.coords) # (lon, lat) tuples
lats = np.array([c[1] for c in coords])
lons = np.array([c[0] for c in coords])
return lats, lons
def _projected_square_cell_vertices(lat_c, lon_c, proj, cell_side_km=100,
n_edge=40):
"""Square cell of side `cell_side_km` in projected space of `proj`,
centred on the projection of (lat_c, lon_c). Returns (lats, lons).
"""
side_m = cell_side_km * 1000.0
half = side_m / 2.0
xy = proj.transform_point(float(lon_c), float(lat_c), ccrs.PlateCarree())
cx, cy = xy[0], xy[1]
# Densify the square's edges in projected space
s = np.linspace(-half, half, n_edge)
bottom = [(cx + dx, cy - half) for dx in s]
right = [(cx + half, cy + dy) for dy in s]
top = [(cx + dx, cy + half) for dx in s[::-1]]
left = [(cx - half, cy + dy) for dy in s[::-1]]
xs = np.array([p[0] for p in bottom + right + top + left])
ys = np.array([p[1] for p in bottom + right + top + left])
# Unproject every vertex back to (lon, lat)
pc = ccrs.PlateCarree()
pts = pc.transform_points(proj, xs, ys)
lons = pts[:, 0]
lats = pts[:, 1]
return lats, lons
def mollweide_cell_vertices(lat_c, lon_c, cell_side_km=100):
"""Mollweide-projected square cell of given side, unprojected to lat-lon."""
return _projected_square_cell_vertices(
lat_c, lon_c, ccrs.Mollweide(), cell_side_km=cell_side_km,
)
def eea_cell_vertices(lat_c, lon_c, cell_side_km=100):
"""EEA reference grid (LAEA Europe / EPSG:3035) square cell."""
eea_proj = ccrs.LambertAzimuthalEqualArea(
central_longitude=10.0, central_latitude=52.0,
false_easting=4_321_000.0, false_northing=3_210_000.0,
)
return _projected_square_cell_vertices(
lat_c, lon_c, eea_proj, cell_side_km=cell_side_km,
)Aspect-ratio and area metrics¶
For each cell we compute its true geographic extent in km and report the aspect ratio (longest / shortest principal axis). Aspect ratio = 1 means a compact, square-ish neighborhood; aspect ratio = 6 means the cell is six times wider than tall.
def laea_xy(lats, lons, lat0, lon0):
"""Lambert azimuthal equal-area projection centred at (lat0, lon0)."""
lat_r = lats * DEG
lon_r = lons * DEG
lat0_r = lat0 * DEG
lon0_r = lon0 * DEG
cosc = (np.sin(lat0_r) * np.sin(lat_r)
+ np.cos(lat0_r) * np.cos(lat_r) * np.cos(lon_r - lon0_r))
k = np.sqrt(2 / (1 + cosc))
x = R * k * np.cos(lat_r) * np.sin(lon_r - lon0_r)
y = R * k * (np.cos(lat0_r) * np.sin(lat_r)
- np.sin(lat0_r) * np.cos(lat_r) * np.cos(lon_r - lon0_r))
return x, y
def cell_metrics(lats, lons, lat_c, lon_c):
"""Width (km), height (km), aspect ratio, area (km²) via LAEA."""
x, y = laea_xy(lats, lons, lat_c, lon_c)
width = x.max() - x.min()
height = y.max() - y.min()
aspect = max(width, height) / min(width, height)
# shoelace area
area = 0.5 * abs(np.dot(x, np.roll(y, 1)) - np.dot(y, np.roll(x, 1)))
return width, height, aspect, areaBuild the figure¶
Three rows of latitudes × three columns of grids. Each tile is a Lambert azimuthal equal-area map centred on the cell, so the cell’s real geographic shape and km-scale extents are visible. The bottom panel plots aspect ratio vs latitude for the three grids over the full biodiversity-relevant range (0–70°N).
Everything is built in a single cell so the figure object stays alive end-to-end (the inline backend auto-closes figures between cells, which would leave the bottom strip detached).
LATITUDES_OF_INTEREST = [0.0, 40.0, 70.0]
NSIDE = 16
# Pre-compute everything for the upper panel grid
grid_specs = [
("Lat-lon 5° × 5°", "tab:red", latlon_cell_vertices),
("Behrmann (equal-area)", "tab:purple",
lambda lc, loc: behrmann_cell_vertices(lc, loc)),
(f"HEALPix DGGS (nside={NSIDE})", "tab:blue",
lambda lc, loc: healpix_cell_vertices(lc, loc, nside=NSIDE)),
]
fig = plt.figure(figsize=(13, 12))
gs = fig.add_gridspec(
4, 3, height_ratios=[1, 1, 1, 1.1], hspace=0.35, wspace=0.20
)
for row, lat_c in enumerate(LATITUDES_OF_INTEREST):
for col, (name, color, builder) in enumerate(grid_specs):
ax = fig.add_subplot(
gs[row, col],
projection=ccrs.LambertAzimuthalEqualArea(
central_latitude=lat_c, central_longitude=0.0
),
)
# Choose a display window large enough for the worst (Behrmann
# at 70°N) cell.
half_extent_km = 1500e3
ax.set_extent(
[-half_extent_km, half_extent_km, -half_extent_km, half_extent_km],
crs=ccrs.LambertAzimuthalEqualArea(
central_latitude=lat_c, central_longitude=0.0
),
)
ax.coastlines(linewidth=0.4, color="0.55")
ax.gridlines(linewidth=0.3, color="0.8", alpha=0.7)
cell_lats, cell_lons = builder(lat_c, 0.0)
ax.fill(
cell_lons, cell_lats,
transform=ccrs.PlateCarree(),
facecolor=color, alpha=0.45, edgecolor=color, linewidth=1.5,
)
w_km, h_km, aspect, area_km2 = cell_metrics(
cell_lats, cell_lons, lat_c, 0.0
)
if row == 0:
ax.set_title(name, fontsize=11, fontweight="bold")
ax.text(
0.02, 0.98,
f"lat = {lat_c:.0f}°N\n"
f"{w_km:.0f} × {h_km:.0f} km\n"
f"aspect = {aspect:.1f}\n"
f"area = {area_km2/1000:.0f} × 10³ km²",
transform=ax.transAxes,
ha="left", va="top",
fontsize=9,
family="monospace",
bbox=dict(facecolor="white", edgecolor="0.7",
boxstyle="round,pad=0.3", alpha=0.85),
)
# --- Aspect ratio across all biodiversity-relevant latitudes ---
# Compares all seven grids in the same plot, grouped by family:
# - Lat-lon (broken)
# - Equal-area projections gridded in projected space (Behrmann, Mollweide, EEA)
# - DGGS family (HEALPix, H3, rHEALPix, ISEA3H)
ax_bottom = fig.add_subplot(gs[3, :])
lat_grid = np.linspace(0, 70, 71)
# Lat-lon (cautionary baseline)
aspect_latlon = [
cell_metrics(*latlon_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
# Projection-based equal-area grids
aspect_behrmann = [
cell_metrics(*behrmann_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
aspect_mollweide = [
cell_metrics(*mollweide_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
aspect_eea = [
cell_metrics(*eea_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
# DGGS family
aspect_healpix = [
cell_metrics(*healpix_cell_vertices(lat, 0.0, nside=NSIDE), lat, 0.0)[2]
for lat in lat_grid
]
aspect_healpix_geo = [
cell_metrics(*healpix_geo_cell_vertices(lat, 0.0, depth=4), lat, 0.0)[2]
for lat in lat_grid
]
aspect_h3 = [
cell_metrics(*h3_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
aspect_rhealpix = [
cell_metrics(*rhealpix_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
aspect_isea3h = [
cell_metrics(*isea3h_cell_vertices(lat, 0.0), lat, 0.0)[2]
for lat in lat_grid
]
# Tier 1 (broken)
ax_bottom.plot(lat_grid, aspect_latlon, color="tab:red", lw=2.2,
label="Lat-lon 5° (broken: not equal-area, not compact)")
# Tier 2 (equal-area projections, gridded in projected space)
ax_bottom.plot(lat_grid, aspect_behrmann, color="tab:purple", lw=1.6,
label="Behrmann (equal-area projection)")
ax_bottom.plot(lat_grid, aspect_mollweide, color="indigo", lw=1.6,
linestyle="--",
label="Mollweide (equal-area projection)")
ax_bottom.plot(lat_grid, aspect_eea, color="orchid", lw=1.6, linestyle=":",
label="EEA reference grid (LAEA Europe)")
# Tier 3 (DGGS family — preserves shape)
ax_bottom.plot(lat_grid, aspect_healpix, color="tab:blue", lw=2.0,
label=f"HEALPix nside={NSIDE} (DGGS, sphere)")
ax_bottom.plot(lat_grid, aspect_healpix_geo, color="royalblue", lw=1.6,
linestyle="--",
label=f"HEALPix-geo nside={NSIDE} (DGGS, WGS84 — GRID4EARTH)")
ax_bottom.plot(lat_grid, aspect_h3, color="tab:cyan", lw=1.6, linestyle="--",
label="H3 res 3 (DGGS, hexagonal)")
ax_bottom.plot(lat_grid, aspect_rhealpix, color="teal", lw=1.6, linestyle=":",
label="rHEALPix res 4 (DGGS, WGS84 cube-projected)")
ax_bottom.plot(lat_grid, aspect_isea3h, color="seagreen", lw=1.6, linestyle="-.",
label="ISEA3H res 8 (DGGS, hex icosahedral)")
ax_bottom.axhline(1.0, color="0.5", linestyle=":", linewidth=0.8)
ax_bottom.set_xlabel("Latitude (°N)")
ax_bottom.set_ylabel("Cell aspect ratio (longer / shorter axis)")
ax_bottom.set_xlim(0, 70)
ax_bottom.set_ylim(
0.9,
max(aspect_latlon + aspect_behrmann + aspect_mollweide + aspect_eea) * 1.05,
)
ax_bottom.legend(loc="upper left", fontsize=8, framealpha=0.95, ncol=2)
ax_bottom.set_title(
"Cell aspect ratio across biodiversity-relevant latitudes — "
"DGGS family stays compact, projection family distorts",
fontsize=11,
)
fig.suptitle(
"Equal-area is necessary but not sufficient for AI-ready biodiversity grids",
fontsize=14, fontweight="bold", y=0.995,
)
plt.savefig("../images/grid_anisotropy.png", dpi=150, bbox_inches="tight")
plt.show()/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
/home/runner/micromamba/envs/dggs-biodiversity-bias/lib/python3.11/site-packages/pyogrio/raw.py:198: RuntimeWarning: driver FlatGeobuf does not support open option DRIVER
return ogr_read(
What the figure shows¶
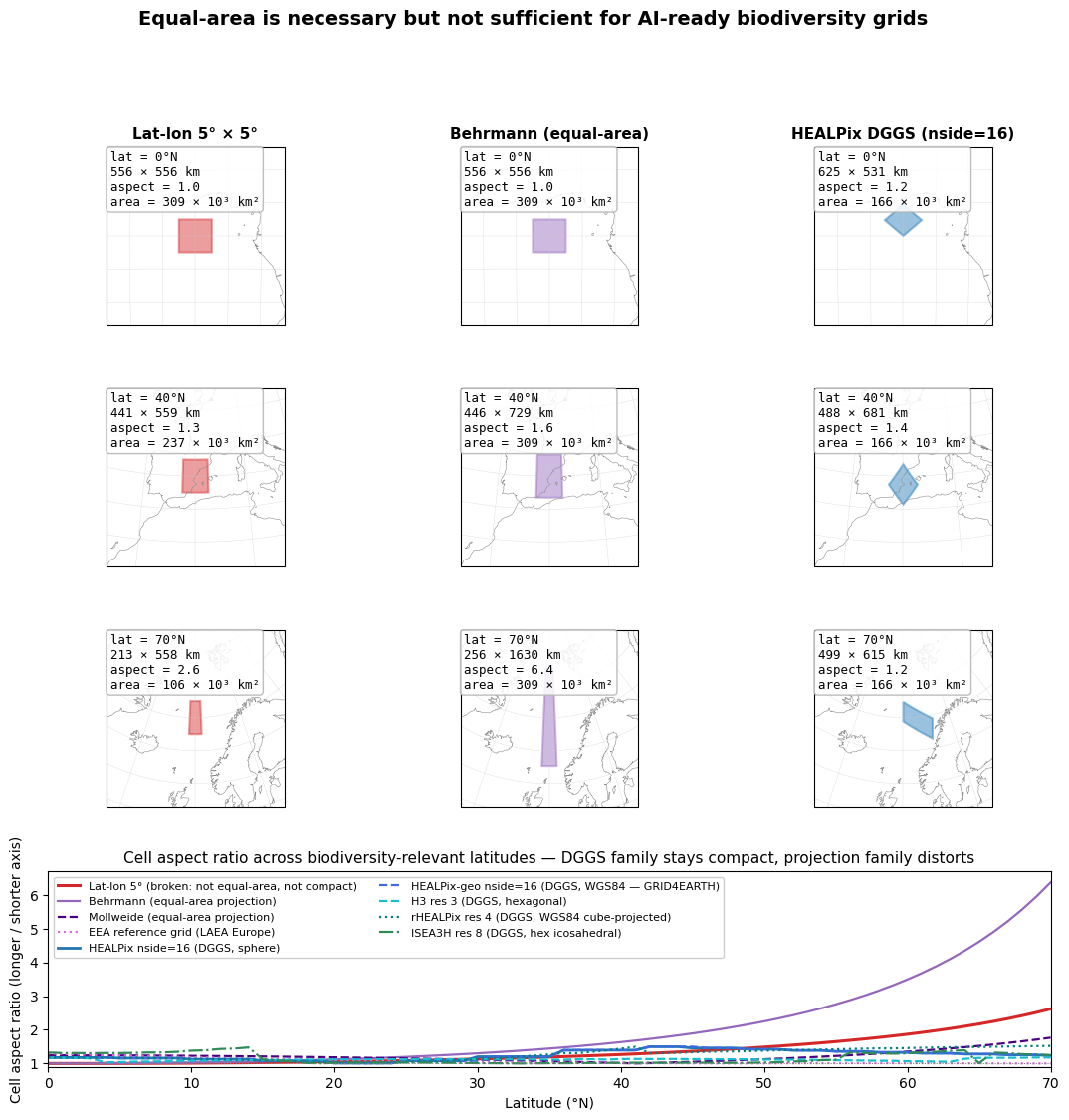
Top three rows. A single cell from each grid system, drawn on a Lambert azimuthal equal-area map centred on the cell. At the equator all three look reasonable. By 40°N the lat-lon cell is already narrower east-west than it is tall, while the Behrmann cell is starting to stretch the other way (wider than tall). At 70°N the anisotropy is dramatic for both rectilinear grids; HEALPix remains compact.
Bottom strip. Aspect ratio (longer axis / shorter axis) as a function of latitude. The dotted line at 1.0 is “perfectly compact.” HEALPix stays close to 1 across the entire 0–70°N band. Lat-lon diverges upward as longitude lines converge. Behrmann diverges upward in the opposite direction — its cells become wider than tall to keep area constant.
The slide-5 takeaway. For mapping and visual presentation, Behrmann or Mollweide are perfectly defensible — they preserve area, which is what global biodiversity counts need. But for machine-actionable workflows — convolutional models, multi- resolution analysis, cloud-native chunked storage — equal-area is necessary but not sufficient. The cell must also be compact (so CNN kernels see consistent neighbourhoods) and hierarchically refinable (so resolution can change without re-projection). HEALPix and other modern DGGSs are designed for both.