Convert EarthCARE MSI Level 2 products to HEALPix at depth 29, following the Sentinel-3 cell-point approach used in legacy-converters.
Key difference from notebook 03: No aggregation. Each MSI pixel maps bijectively to a unique HEALPix cell at depth 29 (~12 mm cell edge). This is a lossless spatial index — every original observation is preserved at its exact location.
| Approach | Depth | Cell size | Aggregation |
|---|---|---|---|
| Notebook 03 (resolution-matched) | 17 | ~458 m | mode/mean/rms |
| This notebook (cell-point) | 29 | ~12 mm | none |
import earthcarekit as eck
import healpix_geo as hpxg
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
import xdggs
from earthcare_dggs.convert import lonlat_to_healpix_cells
from earthcare_dggs.settings import ELLIPSOID
ORBIT = "06109D"
DEPTH = 29 # Cell-point: finest HEALPix level (~12 mm cells)Load MSI products¶
products = {}
for file_type in ["MSI_CM__2A", "MSI_COP_2A", "MSI_AOT_2A"]:
try:
result = eck.search_product(file_type=file_type, orbit_and_frame=ORBIT)
with eck.read_product(result.filepath[0]) as ds:
products[file_type] = ds.load()
print(f"{file_type}: loaded")
except Exception as e:
print(f"{file_type}: not available ({e})")
msi_cm = products["MSI_CM__2A"]
print(f"\nSwath shape: {msi_cm['latitude_swath'].shape}")
print(f"Total pixels: {msi_cm['latitude_swath'].size:,}")MSI_CM__2A: loaded
MSI_COP_2A: loaded
MSI_AOT_2A: loaded
Swath shape: (11216, 357)
Total pixels: 4,004,112
Cell-point mapping at depth 29¶
Each MSI pixel (lon, lat) is assigned to a unique HEALPix cell at depth 29. At this depth, cells are ~12 mm — far smaller than the MSI ~500 m pixel footprint, so each pixel maps to exactly one cell (bijective). No aggregation is needed.
# Extract coordinates
lat = msi_cm["latitude_swath"].values
lon = msi_cm["longitude_swath"].values
valid = np.isfinite(lat) & np.isfinite(lon)
print(f"Valid pixels: {valid.sum():,} / {valid.size:,}")
# Map every valid pixel to a depth-29 cell
cell_ids = lonlat_to_healpix_cells(
lon[valid], lat[valid], depth=DEPTH, ellipsoid=ELLIPSOID,
)
n_unique = len(np.unique(cell_ids))
print(f"\nCell IDs assigned: {len(cell_ids):,}")
print(f"Unique cells: {n_unique:,}")
print(f"Bijective: {n_unique == len(cell_ids)}")Valid pixels: 4,004,112 / 4,004,112
Cell IDs assigned: 4,004,112
Unique cells: 4,004,112
Bijective: True
Build cell-point dataset¶
Unlike notebook 03, we preserve each pixel value as-is, indexed by its depth-29 cell ID. Original dtypes are kept (int8, int16, float32) — no conversion to float.
from earthcare_dggs.convert import detect_fill_value
from earthcare_dggs.settings import MSI_VARIABLES
data_vars = {}
for var_name, config in MSI_VARIABLES.items():
source_ds = products.get(config["source"])
if source_ds is None or var_name not in source_ds:
continue
raw = source_ds[var_name].values.ravel()
# Keep only pixels with valid coordinates
raw_valid = raw[valid.ravel()]
# Mask fill values with NaN (promote to float32 only when fill values exist)
fill = config.get("fill") or detect_fill_value(raw_valid)
if fill is not None:
out = raw_valid.astype(np.float32)
out[raw_valid == fill] = np.nan
elif raw_valid.dtype in (np.float32, np.float64):
out = raw_valid.astype(np.float32)
out[~np.isfinite(raw_valid) | (raw_valid <= -1e30)] = np.nan
else:
out = raw_valid.astype(np.float32)
data_vars[var_name] = out
n_valid = np.isfinite(out).sum()
print(f" {var_name}: {n_valid:,} valid / {len(out):,} pixels")
print(f"\nTotal variables: {len(data_vars)}") cloud_mask: 3,987,502 valid / 4,004,112 pixels
cloud_type: 3,981,555 valid / 4,004,112 pixels
cloud_phase: 954,386 valid / 4,004,112 pixels
surface_classification: 4,004,112 valid / 4,004,112 pixels
quality_status: 4,004,112 valid / 4,004,112 pixels
cloud_top_height: 741,865 valid / 4,004,112 pixels
cloud_top_temperature: 741,865 valid / 4,004,112 pixels
cloud_top_pressure: 741,865 valid / 4,004,112 pixels
cloud_optical_thickness: 748,392 valid / 4,004,112 pixels
cloud_effective_radius: 748,392 valid / 4,004,112 pixels
cloud_water_path: 748,392 valid / 4,004,112 pixels
cloud_top_temperature_error: 741,865 valid / 4,004,112 pixels
isccp_cloud_type: 4,004,112 valid / 4,004,112 pixels
aerosol_optical_thickness_670nm: 684,230 valid / 4,004,112 pixels
aerosol_optical_thickness_865nm: 341,654 valid / 4,004,112 pixels
angstrom_parameter_670nm_865nm: 341,603 valid / 4,004,112 pixels
aerosol_optical_thickness_670nm_error: 684,230 valid / 4,004,112 pixels
aerosol_optical_thickness_865nm_error: 341,654 valid / 4,004,112 pixels
Total variables: 18
# Build xarray Dataset indexed by depth-29 cell IDs
msi_cellpoint = xr.Dataset(
{name: ("cell_ids", data) for name, data in data_vars.items()},
coords={"cell_ids": cell_ids},
)
msi_cellpoint.attrs.update({
"source": "EarthCARE MSI L2A",
"healpix_level": DEPTH,
"healpix_indexing": "nested",
"healpix_ellipsoid": ELLIPSOID,
"orbit_and_frame": ORBIT,
"resampler": "cell-point (no aggregation)",
})
print(msi_cellpoint)<xarray.Dataset> Size: 320MB
Dimensions: (cell_ids: 4004112)
Coordinates:
* cell_ids (cell_ids) uint64 32MB 26284203768...
Data variables: (12/18)
cloud_mask (cell_ids) float32 16MB 3.0 ... nan
cloud_type (cell_ids) float32 16MB 7.0 ... nan
cloud_phase (cell_ids) float32 16MB 1.0 ... nan
surface_classification (cell_ids) float32 16MB -3.276e+04...
quality_status (cell_ids) float32 16MB 0.0 ... 3.0
cloud_top_height (cell_ids) float32 16MB 5.634e+03 ...
... ...
isccp_cloud_type (cell_ids) float32 16MB 8.0 ... -1...
aerosol_optical_thickness_670nm (cell_ids) float32 16MB nan ... nan
aerosol_optical_thickness_865nm (cell_ids) float32 16MB nan ... nan
angstrom_parameter_670nm_865nm (cell_ids) float32 16MB nan ... nan
aerosol_optical_thickness_670nm_error (cell_ids) float32 16MB nan ... nan
aerosol_optical_thickness_865nm_error (cell_ids) float32 16MB nan ... nan
Attributes:
source: EarthCARE MSI L2A
healpix_level: 29
healpix_indexing: nested
healpix_ellipsoid: WGS84
orbit_and_frame: 06109D
resampler: cell-point (no aggregation)
Comparison: depth 29 vs depth 17¶
Show the size difference between the cell-point (depth 29) and aggregated (depth 17) representations.
n_pixels = len(cell_ids)
# Compare with depth 17 (aggregated)
cell_ids_d17 = lonlat_to_healpix_cells(
lon[valid], lat[valid], depth=17, ellipsoid=ELLIPSOID,
)
n_unique_d17 = len(np.unique(cell_ids_d17))
print(f"{'Approach':<25} {'Depth':>6} {'Cells':>12} {'Cell size':>12}")
print("-" * 60)
print(f"{'Cell-point (this nb)':<25} {DEPTH:>6} {n_pixels:>12,} {'~12 mm':>12}")
print(f"{'Aggregated (nb 03)':<25} {17:>6} {n_unique_d17:>12,} {'~458 m':>12}")
print(f"\nCompression ratio (aggregation): {n_pixels / n_unique_d17:.1f}x fewer cells at depth 17")Approach Depth Cells Cell size
------------------------------------------------------------
Cell-point (this nb) 29 4,004,112 ~12 mm
Aggregated (nb 03) 17 4,004,112 ~458 m
Compression ratio (aggregation): 1.0x fewer cells at depth 17
Multi-resolution access¶
A key advantage of depth 29: we can zoom out to any coarser level on the fly
using healpix_geo.nested.zoom_to. Here we demonstrate coarsening from depth 29
to depth 17 (equivalent to notebook 03) and depth 10 (for visualization).
from earthcare_dggs.convert import aggregate_to_healpix
def coarsen_cellpoint(ds, from_depth, to_depth, variables_config):
"""Coarsen a cell-point dataset to a coarser HEALPix depth with aggregation."""
coarse_ids = hpxg.nested.zoom_to(
ds.cell_ids.values, depth=from_depth, new_depth=to_depth,
)
unique_coarse = np.unique(coarse_ids)
coarse_to_idx = {int(c): i for i, c in enumerate(unique_coarse)}
coarse_data = {}
for var_name in ds.data_vars:
vals = ds[var_name].values
mask = np.isfinite(vals)
if mask.sum() == 0:
continue
group_idx = np.array([coarse_to_idx.get(int(c), -1) for c in coarse_ids[mask]])
keep = group_idx >= 0
agg = variables_config.get(var_name, {}).get("agg", "mean")
coarse_data[var_name] = aggregate_to_healpix(
vals[mask][keep], group_idx[keep], len(unique_coarse), method=agg,
)
return xr.Dataset(
{name: ("cell_ids", data) for name, data in coarse_data.items()},
coords={"cell_ids": unique_coarse},
)
# Coarsen to depth 17 (same as notebook 03)
coarse_d17 = coarsen_cellpoint(msi_cellpoint, DEPTH, 17, MSI_VARIABLES)
coarse_d17 = xdggs.decode(
coarse_d17,
grid_info=xdggs.HealpixInfo(level=17, indexing_scheme="nested"),
)
print(f"Coarsened to depth 17: {len(coarse_d17.cell_ids):,} cells")
# Coarsen to depth 10 (for visualization)
coarse_d10 = coarsen_cellpoint(msi_cellpoint, DEPTH, 10, MSI_VARIABLES)
coarse_d10 = xdggs.decode(
coarse_d10,
grid_info=xdggs.HealpixInfo(level=10, indexing_scheme="nested"),
)
print(f"Coarsened to depth 10: {len(coarse_d10.cell_ids):,} cells")Coarsened to depth 17: 4,004,112 cells
Coarsened to depth 10: 21,659 cells
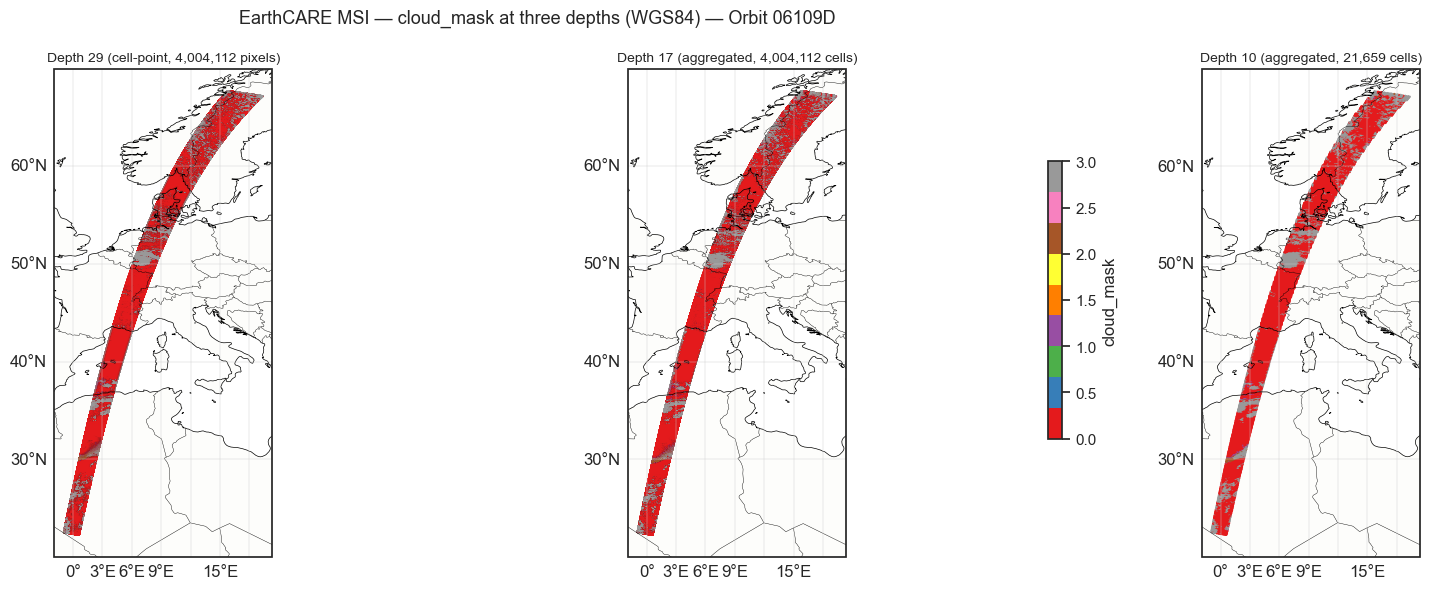
Static visualization¶
Compare the same variable at depth 29 (raw pixels), depth 17, and depth 10.
import cartopy.crs as ccrs
import cartopy.feature as cfeature
var_name = "cloud_mask"
fig, axes = plt.subplots(
1, 3, figsize=(18, 6),
subplot_kw={"projection": ccrs.PlateCarree()},
)
panels = [
(axes[0], msi_cellpoint, DEPTH, f"Depth {DEPTH} (cell-point, {len(cell_ids):,} pixels)"),
(axes[1], coarse_d17, 17, f"Depth 17 (aggregated, {len(coarse_d17.cell_ids):,} cells)"),
(axes[2], coarse_d10, 10, f"Depth 10 (aggregated, {len(coarse_d10.cell_ids):,} cells)"),
]
for ax, ds, depth, title in panels:
ax.add_feature(cfeature.COASTLINE, linewidth=0.5)
ax.add_feature(cfeature.BORDERS, linewidth=0.3)
ax.add_feature(cfeature.LAND, alpha=0.1)
gl = ax.gridlines(draw_labels=True, linewidth=0.3)
gl.top_labels = False
gl.right_labels = False
data = ds[var_name].values
mask = np.isfinite(data)
c_lon, c_lat = hpxg.nested.healpix_to_lonlat(
ds.cell_ids.values[mask], depth=depth, ellipsoid=ELLIPSOID,
)
sc = ax.scatter(
c_lon, c_lat, c=data[mask],
s=0.3, cmap="Set1", vmin=0, vmax=3,
transform=ccrs.PlateCarree(),
)
ax.set_title(title, fontsize=10)
plt.colorbar(sc, ax=axes, shrink=0.6, label=var_name)
plt.suptitle(
f"EarthCARE MSI — {var_name} at three depths ({ELLIPSOID}) — Orbit {ORBIT}",
fontsize=13,
)
plt.tight_layout()
plt.show()/var/folders/zf/53jxd5nj2j3dmfjqpx41p24c0000gn/T/ipykernel_57219/135473460.py:44: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
plt.tight_layout()
Interactive visualization¶
coarse_d10.dggs.explore()Save as DGGS-Zarr¶
Save the cell-point dataset to Zarr. At depth 29, the cell IDs serve as a lossless spatial index — any coarser resolution can be derived on the fly.
import zarr
output_path = "earthcare_msi_cellpoint_06109D.zarr"
# Write to Zarr directly (skip xdggs.encode — no DGGSIndex on depth-29 dataset)
msi_cellpoint.to_zarr(output_path, mode="w")
# Add DGGS Zarr convention metadata
dggs_convention = {
"uuid": "7b255807-140c-42ca-97f6-7a1cfecdbc38",
"name": "dggs",
"schema_url": "https://raw.githubusercontent.com/zarr-conventions/dggs/refs/tags/v1/schema.json",
"spec_url": "https://github.com/zarr-conventions/dggs/blob/v1/README.md",
"description": "Discrete Global Grid Systems convention for zarr",
}
dggs_meta = {
"name": "healpix",
"refinement_level": DEPTH,
"indexing_scheme": "nested",
"ellipsoid": {
"name": "wgs84",
"semimajor_axis": 6378137.0,
"inverse_flattening": 298.257223563,
},
"spatial_dimension": "cell_ids",
"coordinate": "cell_ids",
"compression": "none",
}
root = zarr.open_group(output_path, mode="r+")
root.attrs["zarr_conventions"] = [dggs_convention]
root.attrs["dggs"] = dggs_meta
print(f"Saved to {output_path}")
print(f"Cells: {len(msi_cellpoint.cell_ids):,}")
print(f"Variables: {len(msi_cellpoint.data_vars)}")
print(f"\nDGGS metadata: {dggs_meta}")Saved to earthcare_msi_cellpoint_06109D.zarr
Cells: 4,004,112
Variables: 18
DGGS metadata: {'name': 'healpix', 'refinement_level': 29, 'indexing_scheme': 'nested', 'ellipsoid': {'name': 'wgs84', 'semimajor_axis': 6378137.0, 'inverse_flattening': 298.257223563}, 'spatial_dimension': 'cell_ids', 'coordinate': 'cell_ids', 'compression': 'none'}
/Users/annef/Documents/FAIR2Adapt/DGGS/earthcare-dggs/.pixi/envs/default/lib/python3.13/site-packages/zarr/api/asynchronous.py:247: ZarrUserWarning: Consolidated metadata is currently not part in the Zarr format 3 specification. It may not be supported by other zarr implementations and may change in the future.
warnings.warn(
# Verify round-trip: re-open and decode with xdggs
ds_check = xr.open_zarr(output_path)
ds_check = xdggs.decode(
ds_check,
grid_info=xdggs.HealpixInfo(level=DEPTH, indexing_scheme="nested"),
)
print(ds_check)
print(f"\nIndex type: {type(ds_check.xindexes['cell_ids'])}")
print(f"Cell IDs match: {np.array_equal(ds_check.cell_ids.values, msi_cellpoint.cell_ids.values)}")<xarray.Dataset> Size: 320MB
Dimensions: (cell_ids: 4004112)
Coordinates:
* cell_ids (cell_ids) uint64 32MB 26284203768...
Data variables: (12/18)
aerosol_optical_thickness_670nm (cell_ids) float32 16MB ...
aerosol_optical_thickness_670nm_error (cell_ids) float32 16MB ...
aerosol_optical_thickness_865nm (cell_ids) float32 16MB ...
aerosol_optical_thickness_865nm_error (cell_ids) float32 16MB ...
angstrom_parameter_670nm_865nm (cell_ids) float32 16MB ...
cloud_effective_radius (cell_ids) float32 16MB ...
... ...
cloud_top_temperature_error (cell_ids) float32 16MB ...
cloud_type (cell_ids) float32 16MB ...
cloud_water_path (cell_ids) float32 16MB ...
isccp_cloud_type (cell_ids) float32 16MB ...
quality_status (cell_ids) float32 16MB ...
surface_classification (cell_ids) float32 16MB ...
Indexes:
cell_ids HealpixIndex(level=29, indexing_scheme=nested, kind=pandas)
Attributes:
source: EarthCARE MSI L2A
healpix_level: 29
healpix_indexing: nested
healpix_ellipsoid: WGS84
orbit_and_frame: 06109D
resampler: cell-point (no aggregation)
zarr_conventions: [{'uuid': '7b255807-140c-42ca-97f6-7a1cfecdbc38', 'na...
dggs: {'name': 'healpix', 'refinement_level': 29, 'indexing...
Index type: <class 'xdggs.healpix.HealpixIndex'>
Cell IDs match: True