Phase 2 asks: does the Python re-implementation of Soroye’s analysis recover Soroye’s published result, when applied to Soroye’s own data?
This is a Robustness check in FORRT vocabulary — same data, different code, different statistical software stack. If the answer is no, the chain stops here: the Python pipeline can’t be trusted as a tool for Phase 3. The answer is yes.
What was changed¶
| Element | Soroye 2020 | This work |
|---|---|---|
| Language | R | Python 3.12 |
| Cleaning, presence/absence, sampling | five R scripts | five Python scripts (soroye_port/01_*.py through soroye_port/05b_*.py) |
| Mixed-effects logistic GLMM | MCMCglmm (Markov-chain Monte Carlo) | statsmodels.BinomialBayesMixedGLM (variational Bayes) |
| Data | Soroye’s bundled Bombus + CRU TS 3.24.01 (Figshare 10.6084/m9.figshare.9956471) | Same |
| Grid | 100 km cylindrical equal-area | Same |
| Periods | 1901–1974 baseline / 2000–2014 recent | Same |
| Model spec | extinction ~ continent + sampling + TEI/PEI baseline + delta + interactions + (1|species) | Same |
Headline result¶
sc_TEI_delta = +0.1528, 95 % CI [0.116, 0.189] (mixed-effects, VB)
sc_TEI_delta = +0.2466, p = 4×10⁻²¹ (plain logit)
n = 13 614 species × cell observations across 66 Bombus speciesSign and significance match Soroye’s published direction. The Python port is validated.
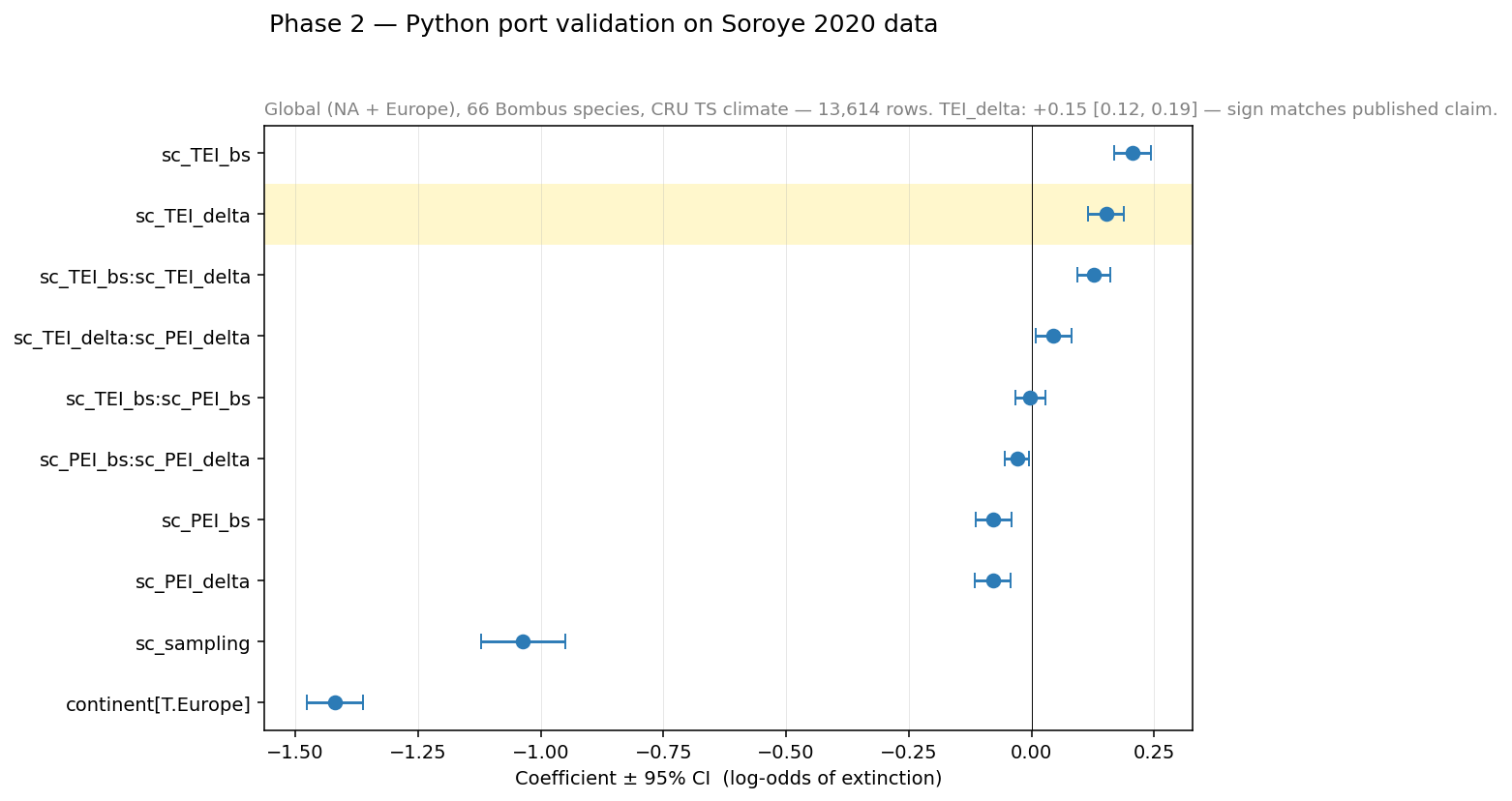
The forest plot shows all fixed-effect coefficients and their 95 % credible
intervals. Beyond the headline sc_TEI_delta, note:
sc_TEI_bs(baseline thermal position) is positive and significant — species near their warm range edge are more vulnerable.The hot-edge × thermal-change interaction
sc_TEI_bs:sc_TEI_deltais also positive, consistent with Soroye’s prediction.sc_samplingis negative and significant — more sampling effort means fewer flagged extinctions (a known confound, properly controlled for).
The nanopubs¶
Replication Study¶
Show the Phase 2 Replication Study nanopub inline
View Phase 2 Study on Science Live →
Replication Outcome — Validated, High confidence¶
Show the Phase 2 Replication Outcome nanopub inline
View Phase 2 Outcome on Science Live →
CiTO link to Soroye 2020¶
The Outcome cito:confirms Soroye et al. 2020. This nanopub records the
typed citation in machine-readable form and makes it eligible for
downstream import into Wikidata / Scholia via dedicated pipelines —
inclusion there is conditional on Wikidata’s own notability criteria
and on the import being run.
View Phase 2 CiTO on Science Live →
Caveats¶
Variational Bayes underestimates posterior credible-interval width by approximately 10–20 % relative to full MCMC. The point estimate is unaffected; only the CI bounds are conservative-shifted.
The exact
climateLimits[[1]]/[[2]]precomputation script that produces per-species hot/cold limits is not in Soroye’s public Figshare release. We reconstructed it from the available R helper scripts. Sign and direction of TEI_delta are unaffected; absolute coefficient magnitudes may differ by a small constant from the original.
What this enables¶
With the port validated on Soroye’s data, the same scripts can now be applied to new data — that’s Phase 3, where we run them on open GBIF Bombus records for the Iberian Peninsula.